HypoEvolve
Benchmarking LLM-Generated Biological Hypotheses for Scientific Discovery
Jefferson Chen, Samuel Lee, Jieyuan Liu, Zhiting Hu, Zhen Wang
University of California, San Diego
jec068@ucsd.edu · hsl023@ucsd.edu · jil029@ucsd.edu · zhh019@ucsd.edu · zhw085@ucsd.edu
Elevator pitch
HypoEvolve is a multi-agent evolutionary system that refines LLM-generated biomedical hypotheses using selection, crossover, and mutation (genetic algorithms).
We show that evolutionary refinement yields substantial gains under external biological validation on drug repurposing (DepMap CRISPR dependency) and improves performance on Type 2 Diabetes (T2D) gene discovery.
Target audience / stakeholder: computational biology researchers, ML-for-science practitioners, and reviewers who need a systematic way to improve (not just generate) mechanistic hypotheses.
Project resources (quick links)
- GitHub repo: https://github.com/JeffersonChen888/HypoEvolve
- Postal (PDF): https://drive.google.com/file/d/1jCgbqp21dvVsrzsLfzA9gf32aaDAURsr/view?usp=sharing
- Final report: https://drive.google.com/file/d/1VlOpv-IeNF-nmLtqVthjJpcdaTL48wv5/view?usp=sharing
Table of Contents
- Key results
- Problem & scope
- Methods
- Experiments & data
- Interpretation
- Limitations & next steps
- References
Key results
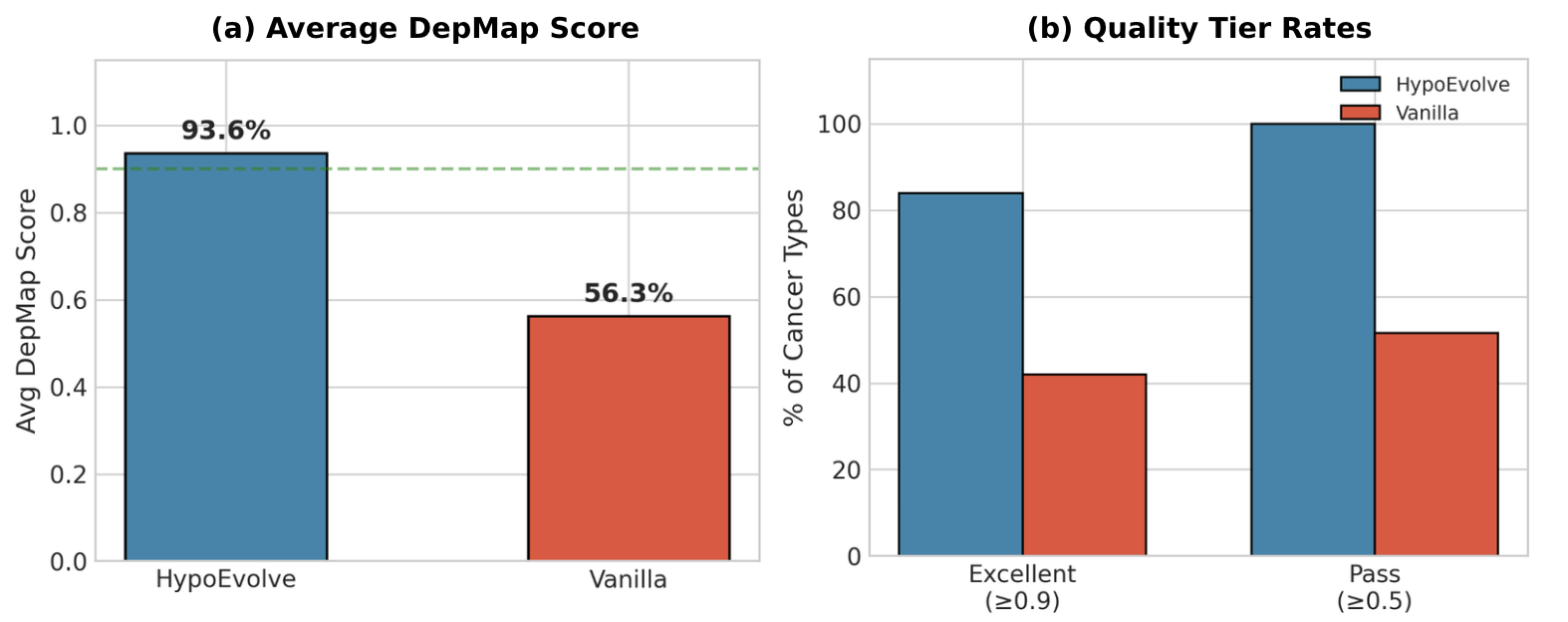 Figure 1. External validation comparison between single-pass prompting and HypoEvolve (drug repurposing).
Figure 1. External validation comparison between single-pass prompting and HypoEvolve (drug repurposing).
What we found:
- Evolutionary refinement improves externally validated hypothesis quality versus single-pass prompting.
- Gains appear across many cancer types, indicating robustness.
- Improvements are achieved without DepMap leakage (DepMap is not used during generation/scoring—only for external evaluation).
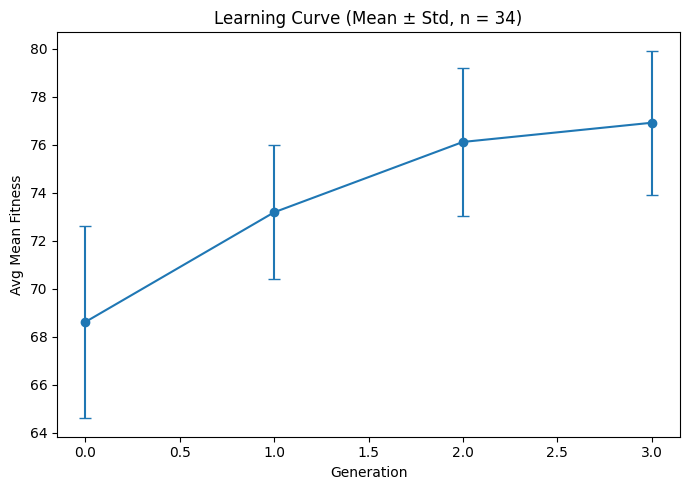 Figure 2. Learning curve showing LLM-evaluated fitness improving and stabilizing across generations (drug repurposing).
Figure 2. Learning curve showing LLM-evaluated fitness improving and stabilizing across generations (drug repurposing).
Problem & scope
Problem
Single-pass LLM prompting produces one hypothesis with no systematic refinement. Unlike researchers, LLM workflows typically lack selection pressure to preserve strong ideas and revise weak ones.
Scope boundaries
We do:
- Treat hypothesis refinement as evolutionary optimization (selection + crossover + mutation).
- Evaluate on two tasks:
1) Drug repurposing with external validation using DepMap CRISPR dependency
2) Type 2 Diabetes (T2D) gene discovery comparisons
We do not:
- Conduct wet-lab validation or claim clinical readiness.
- Fine-tune foundation models.
Methods
System overview
HypoEvolve is a multi-agent GA-style loop:
- Generation Agent: proposes a population of candidate hypotheses
- Reflection Agent: performs structured review and assigns component scores
- Evolution Agent: produces offspring via recombination and mutation
- Supervisor Agent: runs tournament selection + elitism and advances the population
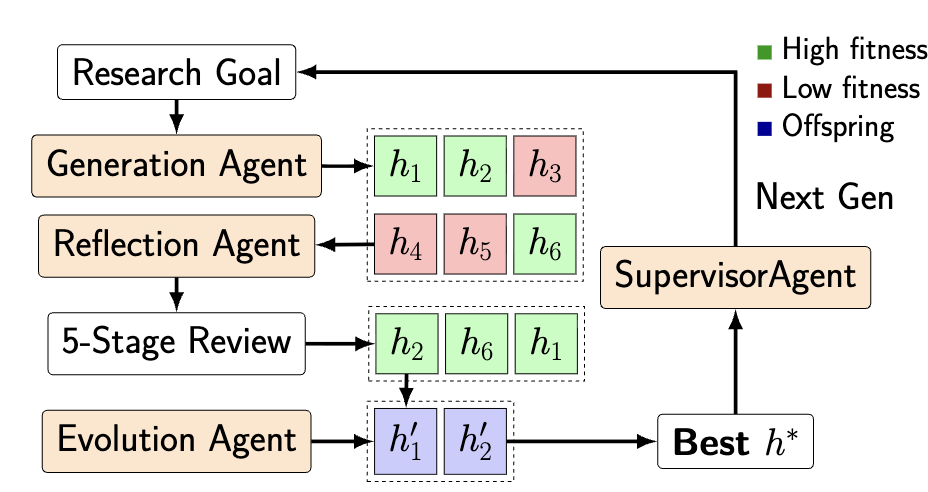 Figure 3. Multi-agent evolutionary loop used to iteratively refine hypotheses.
Figure 3. Multi-agent evolutionary loop used to iteratively refine hypotheses.
Evolutionary cycle per generation
Each generation follows:
- Review: generate component scores
- Selection: tournament selection + elitism
- Crossover: recombine mechanistic reasoning
- Mutation: controlled semantic edits
Objective
We define hypothesis search as:
$h^* = \arg\max_h f(h)$
where fitness is:
$f(h) = w_c s_c + w_n s_n + w_q s_q$
- $s_c$: correctness (biological plausibility)
- $s_n$: novelty (non-trivial / less redundant mechanisms)
- $s_q$: quality (clarity + explanatory strength)
Weights balance novelty and quality while maintaining correctness.
Experiments & data
Task 1: Drug repurposing (oncology)
Given a cancer type, the system proposes:
- an FDA-approved drug
- a pathway / target gene
- a mechanistic rationale
External validation dataset: DepMap CRISPR dependency.
DepMap aggregates genome-scale CRISPR knockout screens across many cancer cell lines. Dependency scores indicate whether a gene is essential for cell survival in a given context. We use DepMap only after hypotheses are generated to measure whether implicated genes show strong dependency in the relevant cancer setting (external validation).
Leakage control: DepMap is not used in internal scoring or during evolution—only for post-hoc evaluation.
Task 2: Type 2 Diabetes (T2D) gene discovery
We also evaluate HypoEvolve on identifying T2D-associated genes, comparing performance across different LLM backbones (as in the poster).
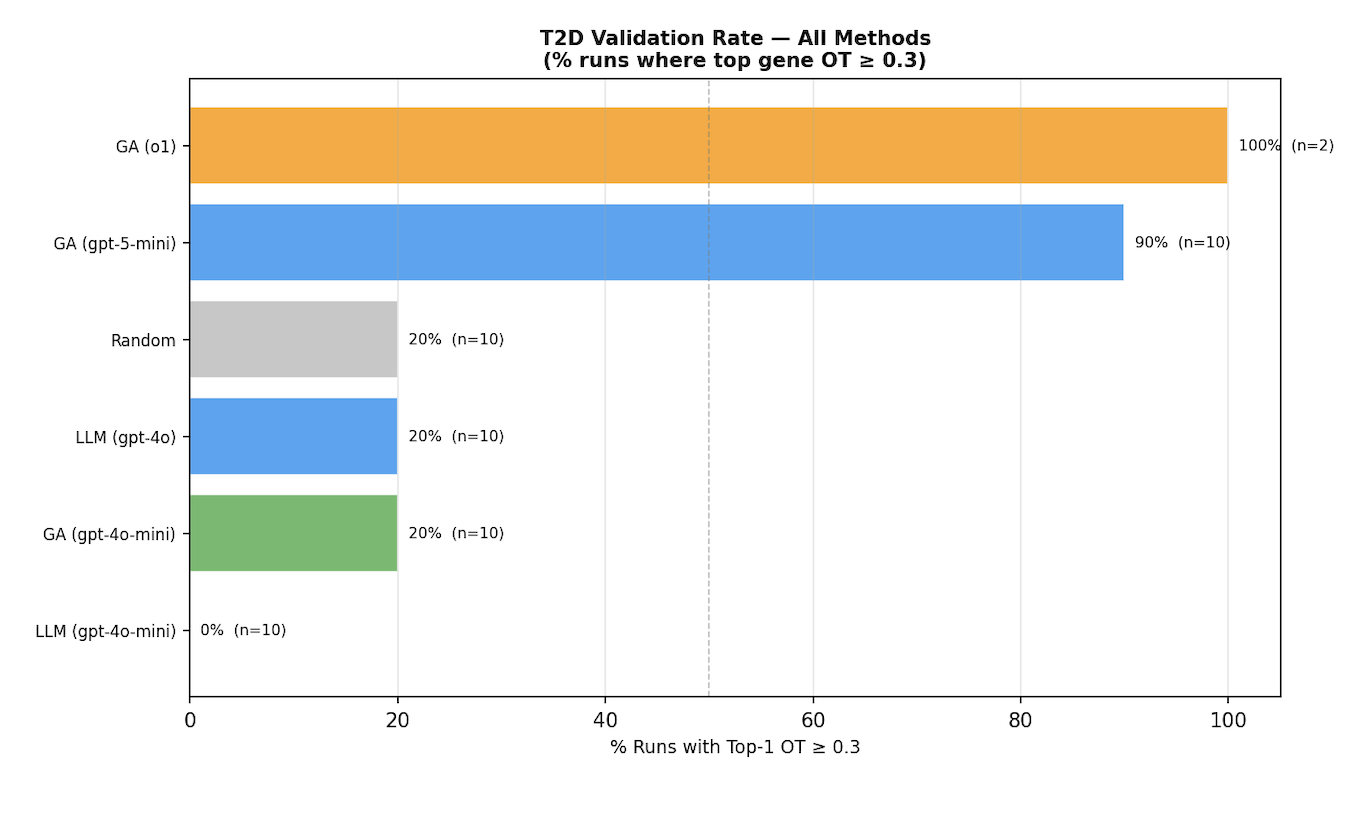 Figure 4. T2D results: single-pass vs HypoEvolve across different LLMs.
Figure 4. T2D results: single-pass vs HypoEvolve across different LLMs.
Interpretation
What changed across generations (qualitative):
- Biologically inconsistent explanations are filtered out early by selection pressure.
- Crossover can combine partial mechanisms into more coherent rationales.
- Mutation often simplifies or repairs brittle reasoning chains.
What to trust vs. what to be cautious about:
- External validation improvements (DepMap) are meaningful because they are evaluated out-of-loop.
- Internal fitness (LLM-based) is useful for steering evolution, but should be treated as a heuristic, not ground truth.
Limitations & next steps
Limitations
- Internal scoring relies on LLM judgments; calibration and human review would strengthen credibility.
- DepMap measures gene dependency, which is an indirect proxy for full mechanistic correctness.
- Compute limits constrain population size and generation depth.
Next steps
- Run ablations: remove crossover, remove mutation, remove elitism.
- Add additional discovery tasks beyond oncology and T2D.
- Incorporate structured biological knowledge (e.g., knowledge graphs) to constrain mutations and improve grounding.
References
[1] Annu Lambora, Kunal Gupta, and Kriti Chopra. Genetic algorithm — a literature review.
In 2019 International Conference on Machine Learning, Big Data, Cloud and Parallel Computing (COMITCon), pp. 380–384. IEEE, 2019.